Leveraging Flywheel for Deep Learning Model Prediction
Since 2012, the Medical Image Computing and Computer Assisted Intervention Society (MICCAI) has put on the Brain Tumor Segmentation (BraTS) challenge with the Center for Biomedical Image Computing and Analytics (CBICA) at the Perelman School of Medicine at the University of Pennsylvania. Past competitions have seen rapid improvements in the automated segmentation of gliomas. This automation promises to address the most labor-intensive process required to accurately assess both the progression and effective treatment of brain tumors.
In this article, we demonstrate the power and potential of coupling the results of this competition with a FAIR (Findable, Accessible, Interoperable, Reusable) framework. With constructing a well-labeled dataset constituting the most labor-intensive component of processing raw data, it is essential to automate this process as much as possible. We utilize Flywheel as our FAIR framework to demonstrate this process.
Flywheel is a FAIR framework that leverages the proprietary core infrastructure with open-source extensions (gears) to collect, curate, compute on, and collaborate on clinical research data. The core infrastructure of a Flywheel instance manages the collection, curation, and collaboration aspects, enabling multi-modal data to be quickly searched across an enterprise-scale collection. Each “gear” of the Flywheel ecosystem is a container-encapsulated open-source algorithm with a standardized interface. This interface enables consistent stand-alone execution or coupling with the Flywheel core infrastructure—complete with provenance of raw data, derived results, and usage records.
For the purposes of this illustration, we wrap into a gear the second-place winner of the MICCAI 2017 BraTS Challenge. This team’s entry is one of the few that has both a docker hub image and a well-documented github repository available. Their algorithm is built around both TensorFlow and NiftyNet frameworks for training and testing their Deep Learning model. As illustrated in our github repository, this “wrapping” constitutes providing the data configuration expected by their algorithm and launching their algorithm for model prediction (*).
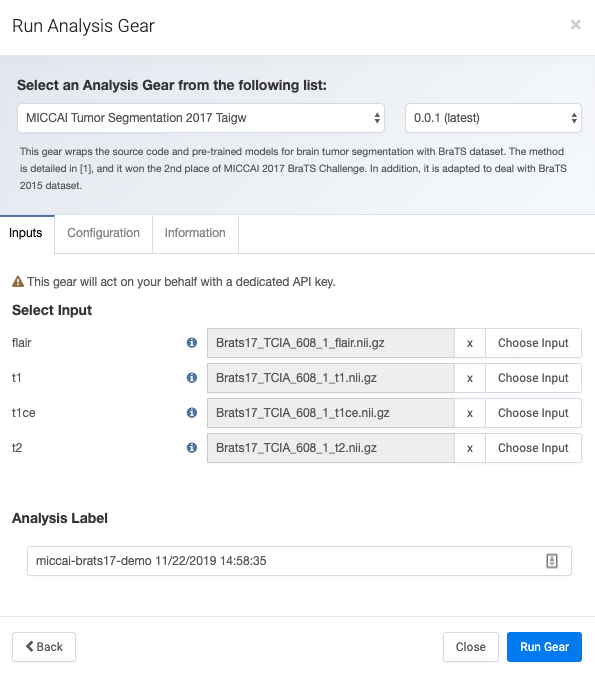
As shown in the figure above, Flywheel provides a user-friendly interface to navigate to the MRI images expected for execution. With the required co-registered and skull-stripped MRI modalities (T1-weighted, T1-weighted with contrast, T2-weighted, and Fluid Attenuation Inversion Recovery), segmentation into distinct tissues (normal, edema, contrast enhancing, and necrosis) takes twelve minutes on our team’s Flywheel instance (see figure below). This task can take a person over an hour to segment the same tumor. When performed on a Graphical Processing Unit (GPU), this task takes less than three minutes to complete.

Segmentation into normal, edema, contrast enhancing, and necrosis tissues with the Flywheel-wrapped second place winner of the 2017 BraTS Challenge.
Although this example predictively segments the tumor of a single patient, modifications to this gear can allow tumor segmentation of multiple patients for multiple imaging sessions over the course of their care. Furthermore, with scalable cloud architecture, these tasks can be deployed in parallel, significantly reducing the overall time required to iterate inference over an entire image repository. Enacting this as a pre-curation strategy could significantly reduce the time necessary for manual labeling of clinical imaging data.
Therein lies the vast potential benefit from using a strong FAIR framework in an AI-mediated workflow. Being able to pre-curate new data, optimize human input, and retrain on well-labeled data over accelerated time-scales. These model design, train, and test cycles are greatly facilitated by a FAIR framework, which is able to curate the data, results, and their provenance in a searchable interface.
As with this brain tumor challenge example, there are many other similar challenge events that make their algorithms and pretrained models publicly available for the research community. One nexus of these is the Grand Challenges in Biomedical Image Analysis, hosting over 21,000 submissions in 179 challenges (56 public, 123 hidden). Flywheel’s capacity to quickly package these algorithms to be interoperable with its framework makes it a powerful foundation for a data-driven research enterprise.
Two more useful deep learning and GPU-enabled algorithms have recently been incorporated into Flywheel gears. First, quickNAT uses default or user-supplied pre-trained deep learning models to segment neuroanatomy within thirty seconds when deployed on sufficient GPU hardware. We have wrapped a Pytorch implementation of quickNAT in a Flywheel gear. Prediction of brain regions on CPU hardware requires two hours. Although much longer than thirty seconds needed on a GPU, it is still a fraction of the nearly twelve hours needed for FreeSurfer’s recon-all. Next, we have Nobrainer, a deep learning framework for 3D image processing. The derived Flywheel gear uses a default (or user-supplied) pre-trained model to create a whole brain mask within two minutes on a CPU. Utilizing a GPU brings this time down under thirty seconds.
The previous paragraph elicits two questions. First, with GPU model prediction times significantly faster than CPUs, when will GPU-enabled Flywheel instances be available? The next being, how can Flywheel be effectively leveraged in training deep learning models? Flywheel is actively developing GPU-deployable gears and the architecture to deliver them. We briefly explore the second question next, leaving a more thorough investigation for another article.
Training on an extensive and diverse dataset is needed for Deep Learning models to generalize effectively and accurately across unseen data. With uncommon conditions, such as gliomas, finding enough high-quality data at a single institution can be daunting. Furthermore, sharing these data across institutional boundaries incurs the risk of exposing protected health information (PHI). With Federated Training, Deep Learning models (and their updates) are communicated across institutional boundaries to acquire the abstracted insight of distributed annotation. This eliminates the risk and requirement of transferring large data repositories while still allowing model access to a diverse dataset. With Federated Search across institutional instances of Flywheel firmly on the roadmap, this type of Federated Training of Deep Learning models will be possible within the Flywheel ecosystem.
(*) The authors of this repository and the University College London do not explicitly promote or endorse the use of Flywheel as a FAIR framework.